Single-cell data sets are exploding in size. From a few thousand cells, we now see more than 1 million data points being generated (Cao et. al., 2019). Large single-cell atlases are valuable resources for exploration and insights, but also pose a great challenge to anyone who wants to view and reanalyze them. How can we analyze these huge and complex data sets if our computational capacity falls behind such an exponential increase in the number of data points?
We introduce BioTuring Big Data Portal, a new gateway that allows scientists worldwide to connect to our server and explore our curated single-cell transcriptome database of 6.5 million cells from more than 150 latest publications – without any computational limits.
By letting scientists perform heavy analyses on BioTuring server, the Portal for the first time enables interactive visualizations and real-time analysis on huge single-cell data sets on a normal laptop. This came from a lot of engineering effort: 100X in speed improvement of gene marker finding algorithm, data compression, improvement in sparse matrix read/write, scatter plot visualization, etc.
To aid global research during the COVID-19 lockdown, access to BioTuring Big Data Portal and our entire single-cell database will be completely FREE for everyone in this month.
To get installation instructions, please visit: https://bioturing.com/bigdata
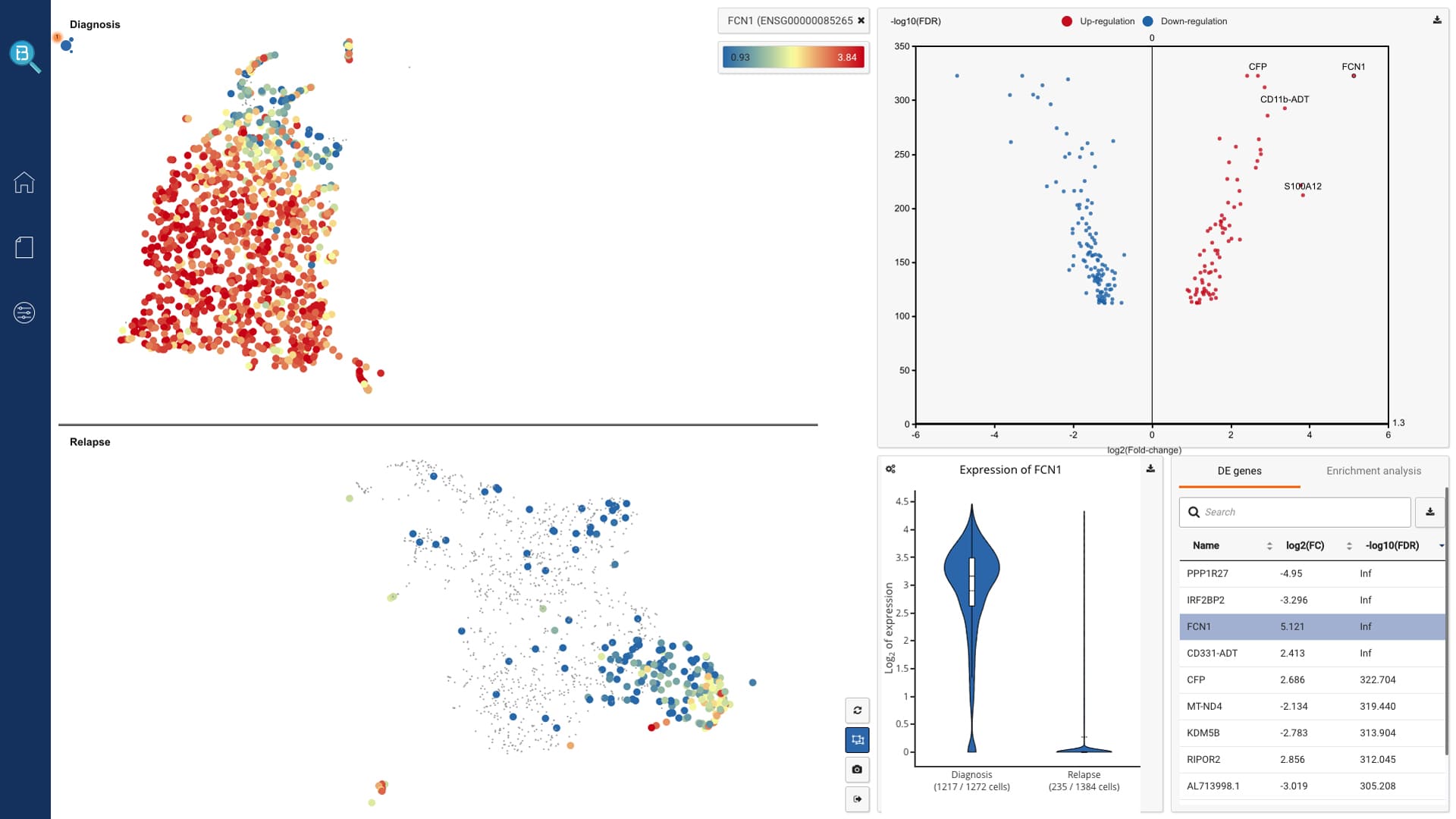
Supported visualizations and analytics
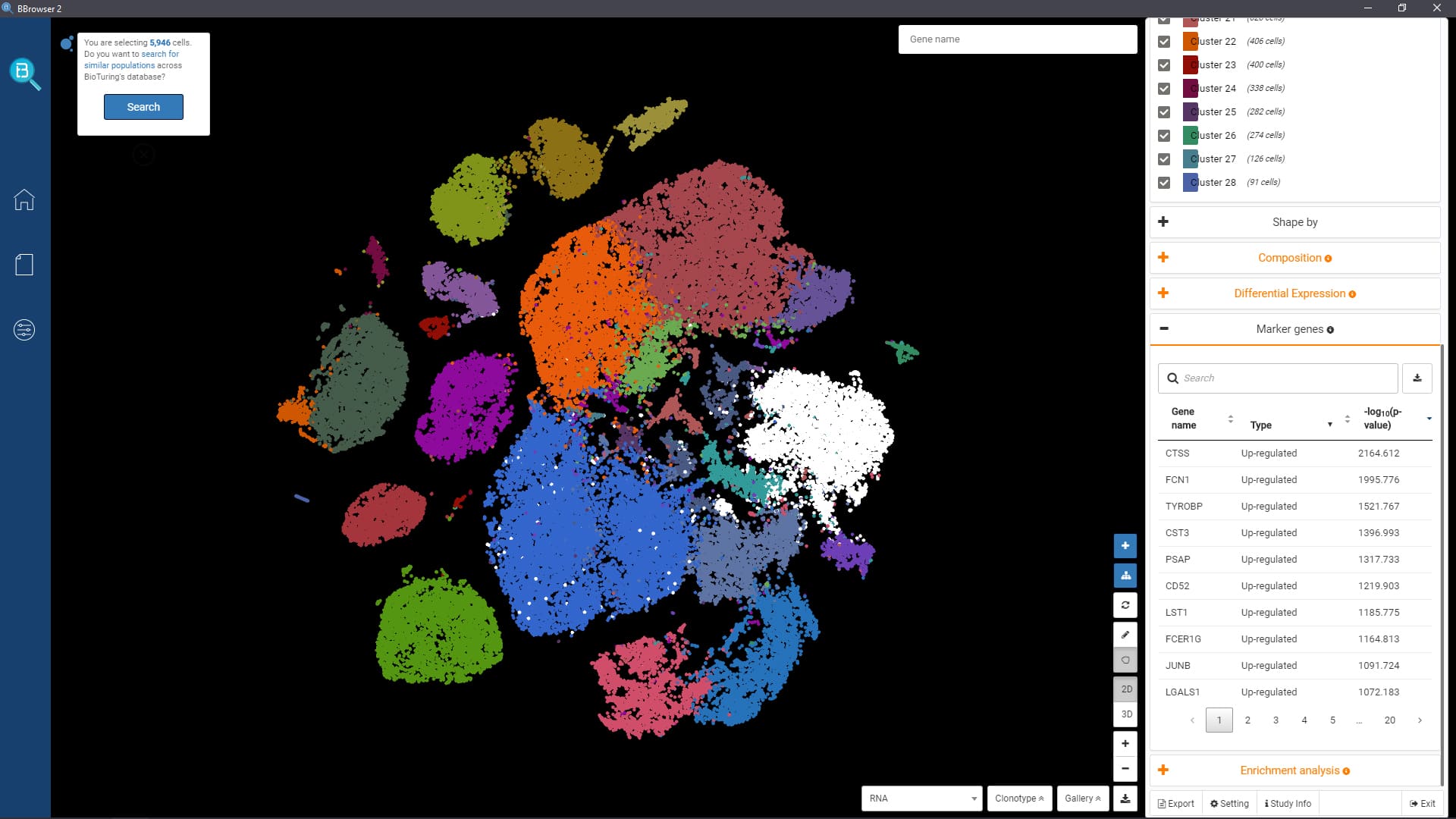
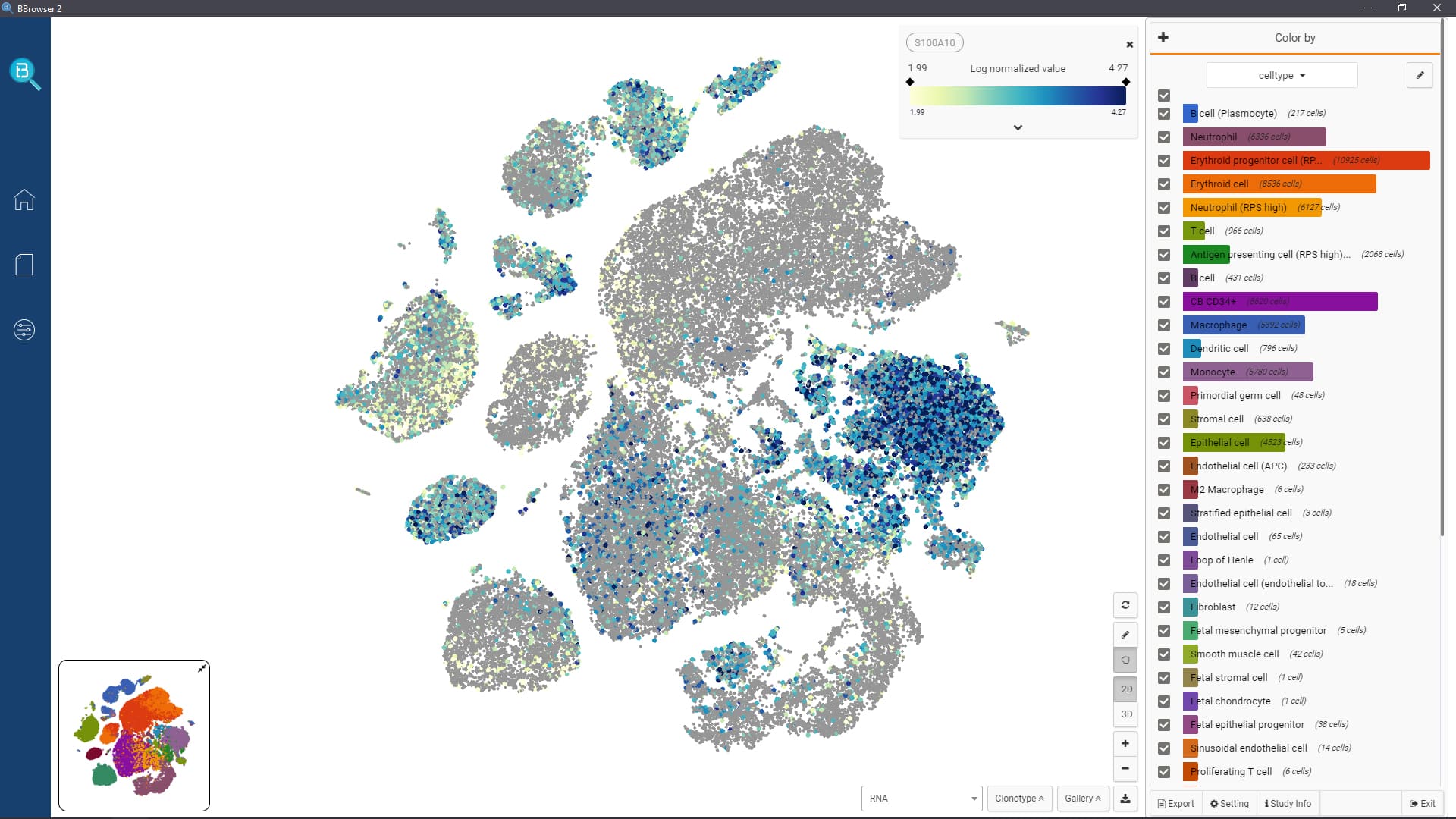
- Interactively visualize public data in 2D or 3D, fully annotated with cell types and metadata
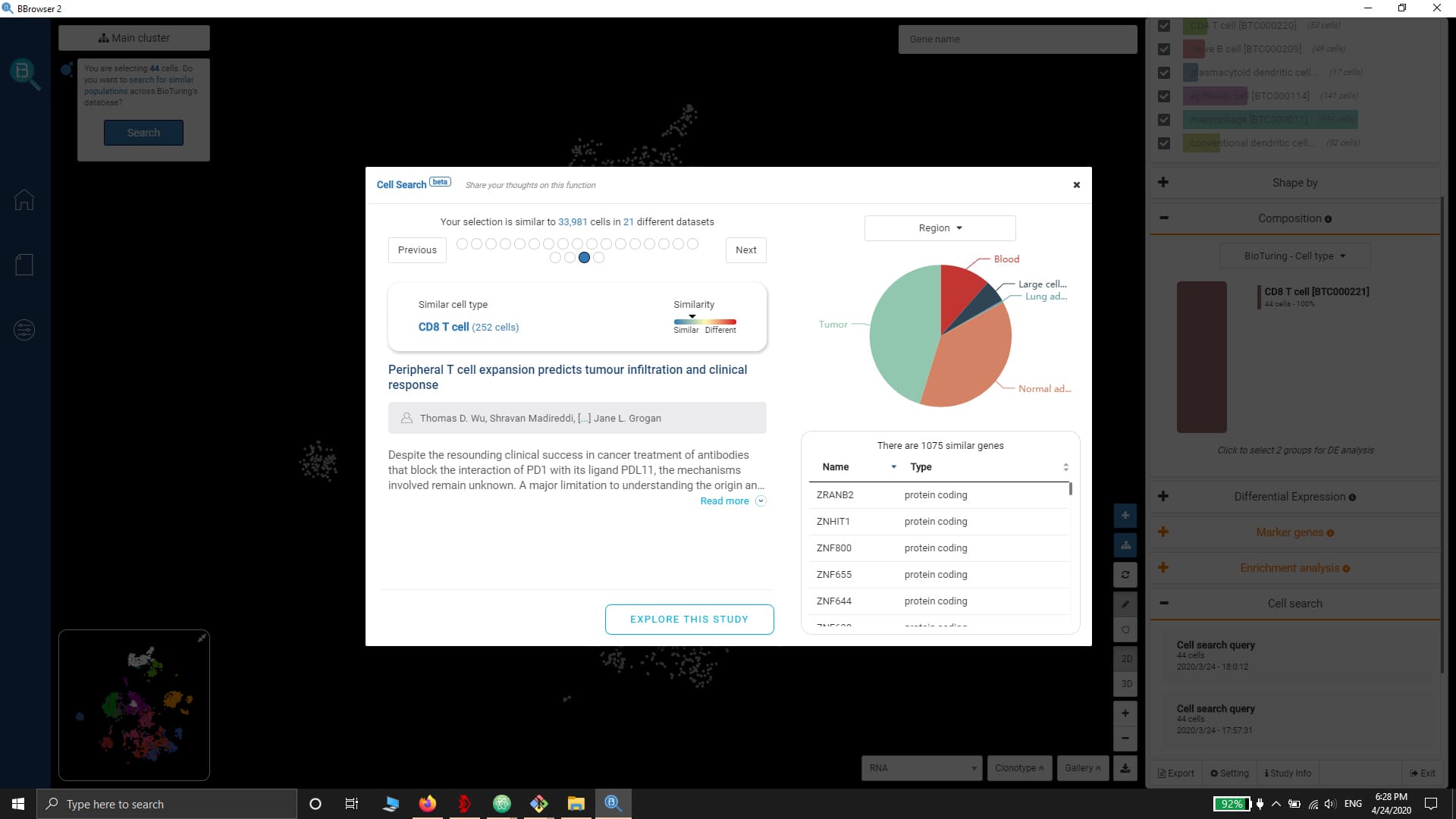
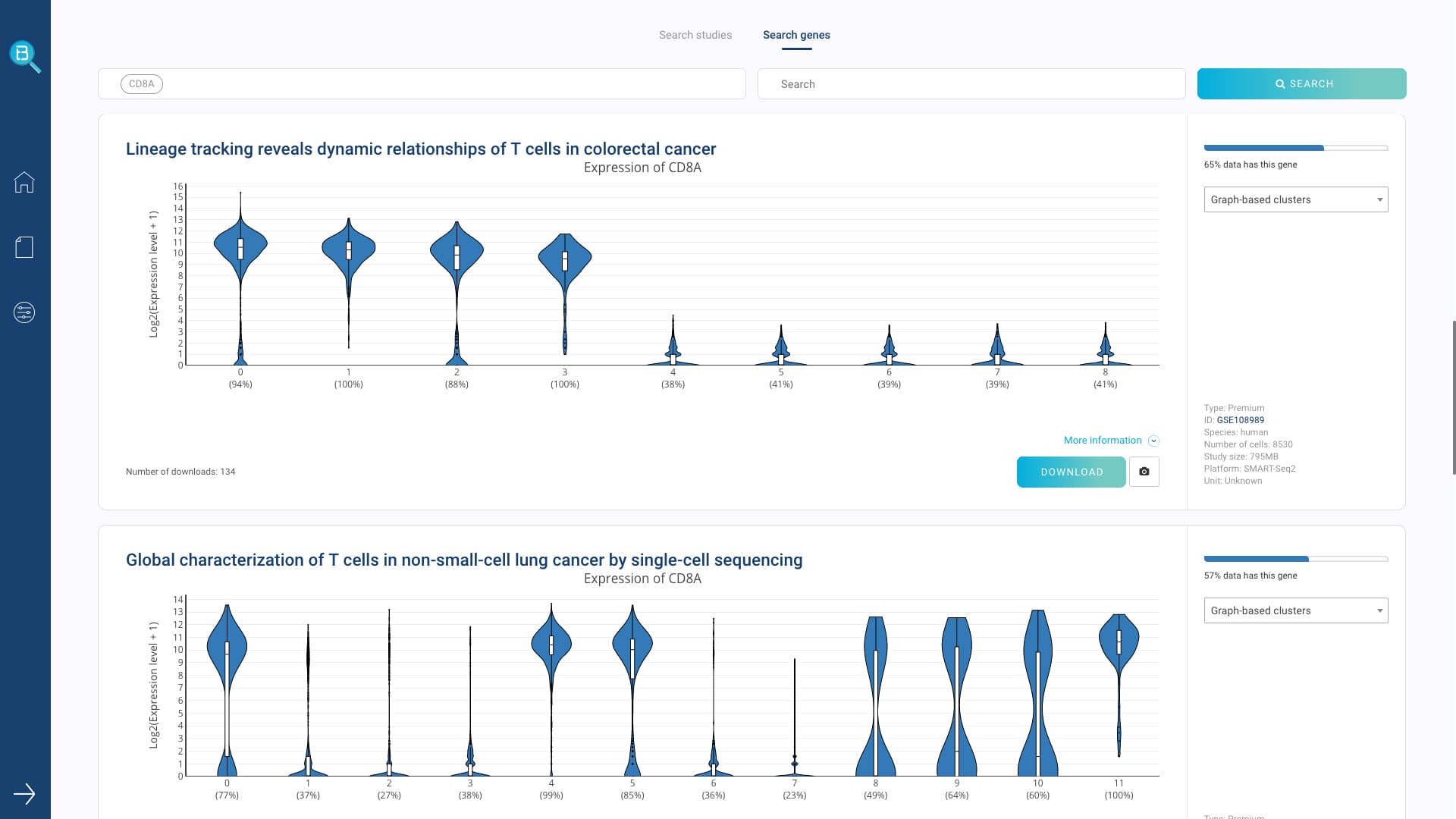
- Query gene expression and search similar cell populations across the database
- Find markers, run enrichment analysis, sub-clustering
- View one or multiple gene expression in the data
- Study composition
- Find differentially expressed genes
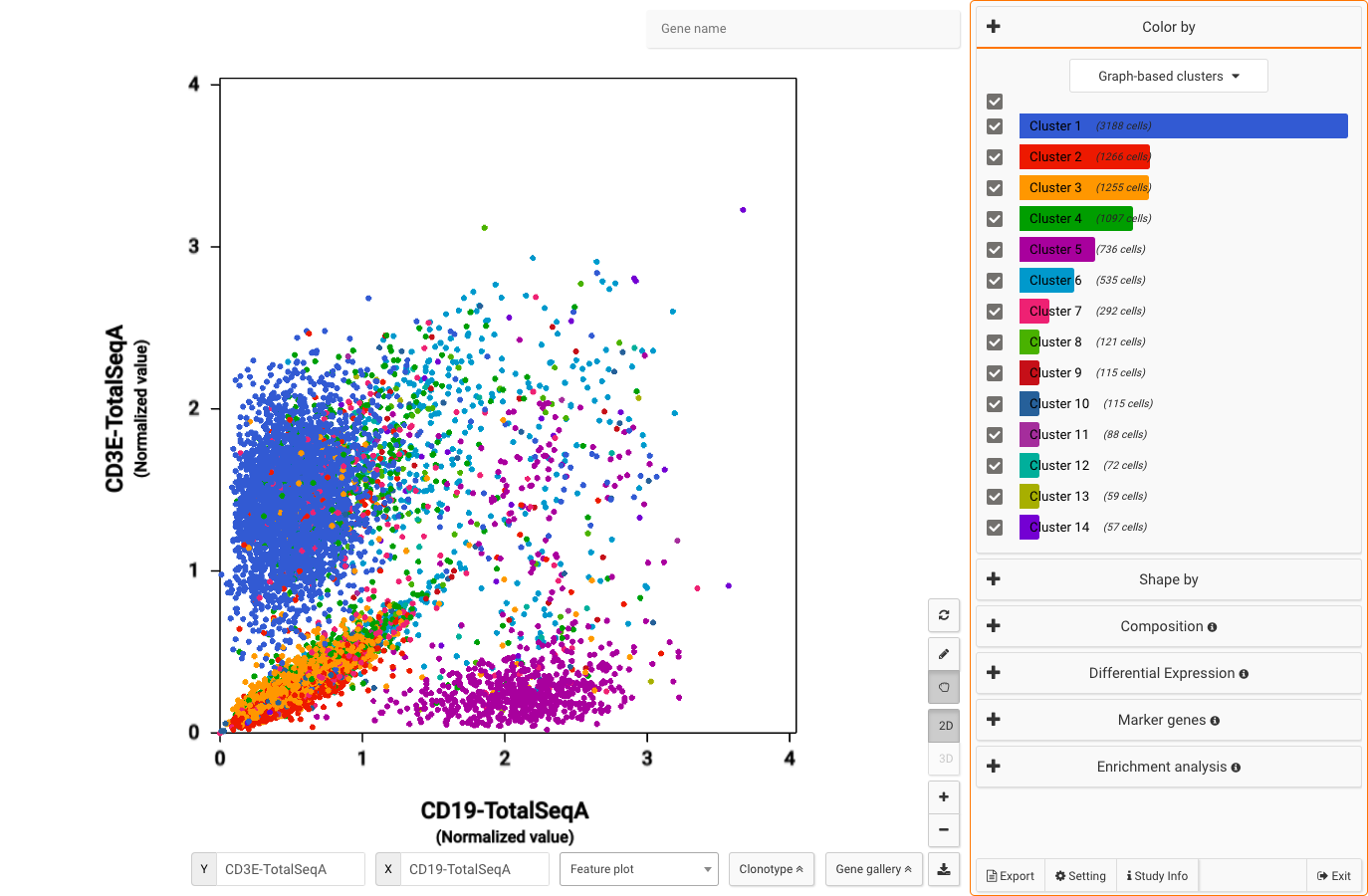
- CITE-Seq and clonotype data analysis
Available data sets
BioTuring Big Data Portal is now hosting more than 6.5 million cells from more than 150 publications, including extra large data sets with more than 100k cells, or even exceeding 1.3 million.
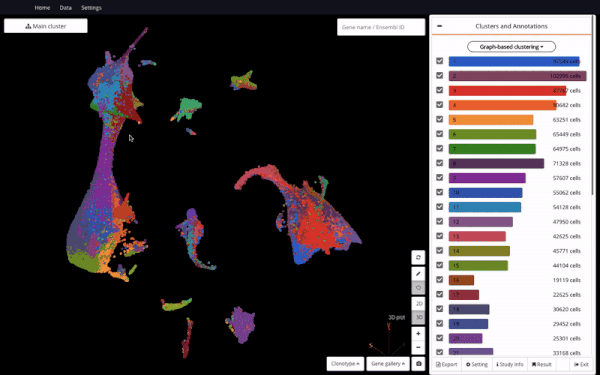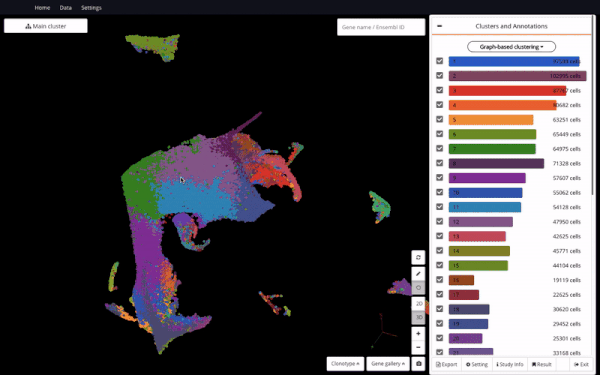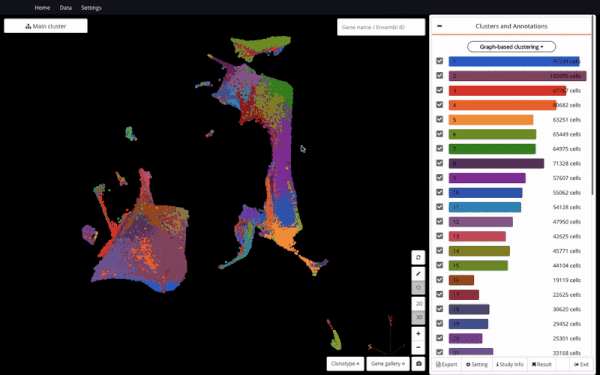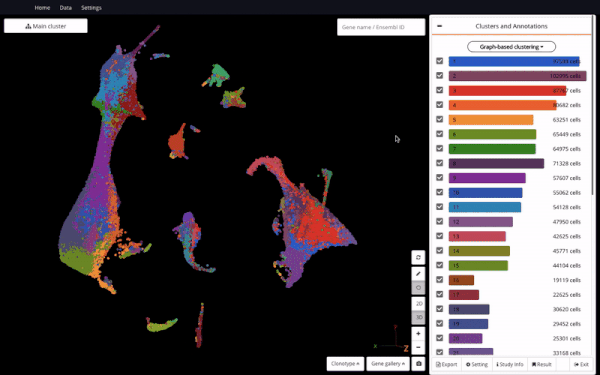
3D UMAP on “The single-cell transcriptional landscape of mammalian organogenesis” (Cao et. al., 2019) (1,331,984 cells)
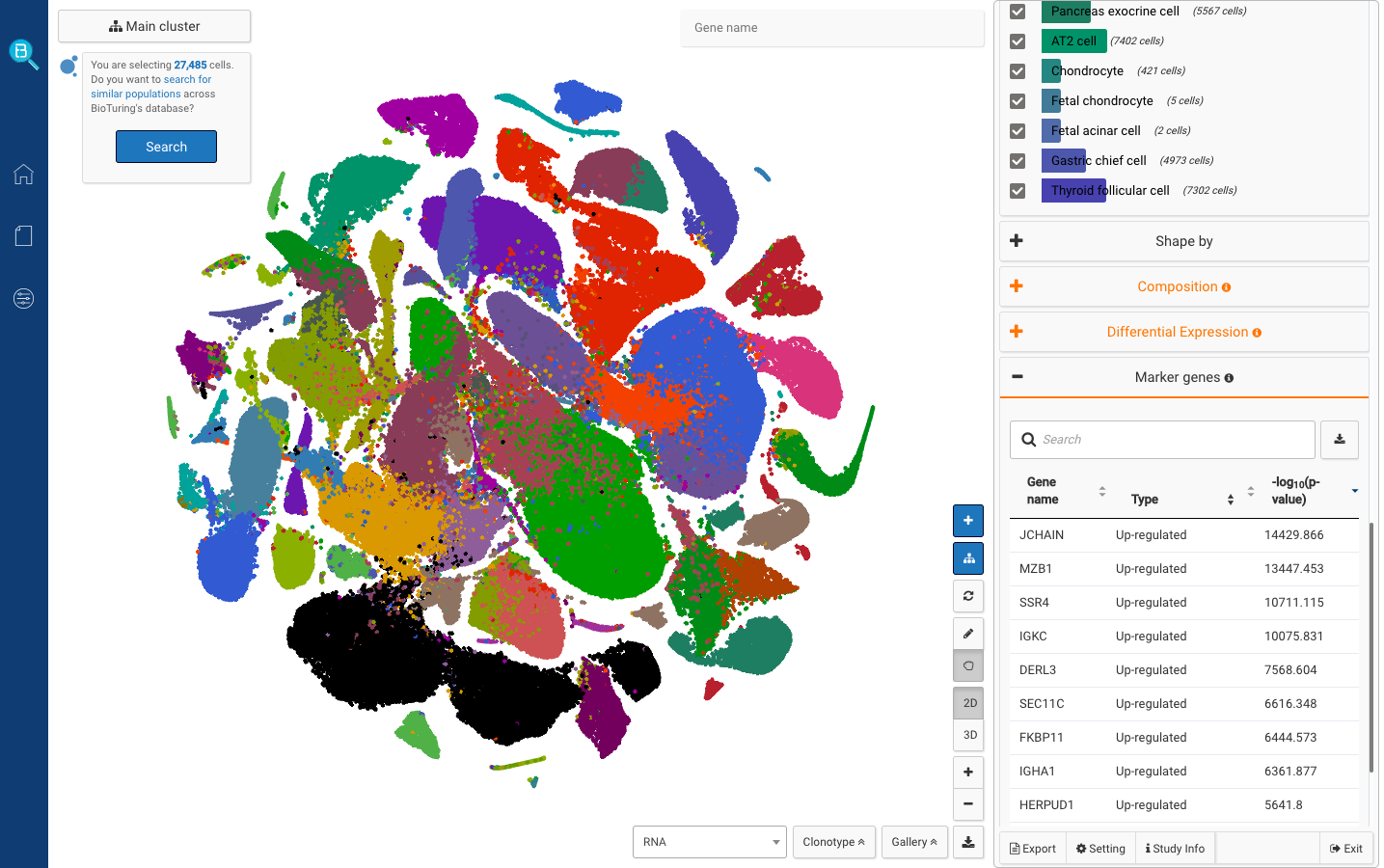
Construction of a human cell landscape at single-cell level (Han et. al., 2020) (Adult samples: 332,577 cells)
To get a full list of available data sets, please visit: https://bioturing.com/bbrowser/datasets
Installation guide: https://bioturing.com/bigdata