Single-cell RNA-seq technologies have opened up a completely new era for transcriptomic studies. For the first time ever, scientists can look at individual transcriptomic profiles of millions of cells, and better understand how each cell functions in a tissue. Yet science is confronting bigger challenges analyzing these massive amounts of data from single-cell RNA-seq: finding the best algorithms to quickly and accurately process such data, and a visualization platform that is strong enough to interactively graph hundreds of thousands to millions of data points.
We introduce BioTuring Browser, the software to tackle major challenges in single-cell RNA-seq data analysis that is extremely easy to use for any biomedical scientists. BioTuring Browser dashboards are interactive, compact enough to run on a standard laptop, and most importantly, has the power to process and visualize up to 1.3 million cells at a time.
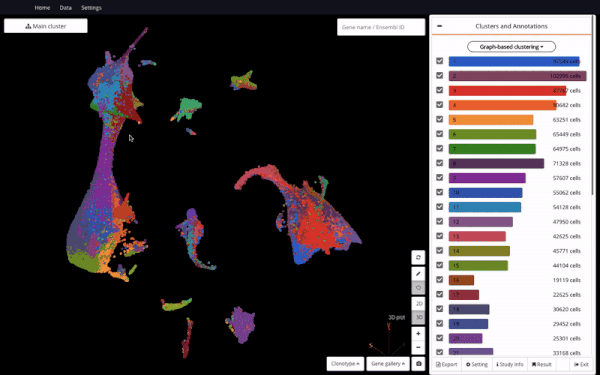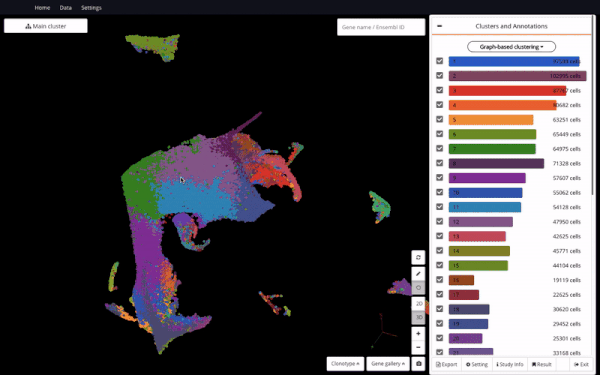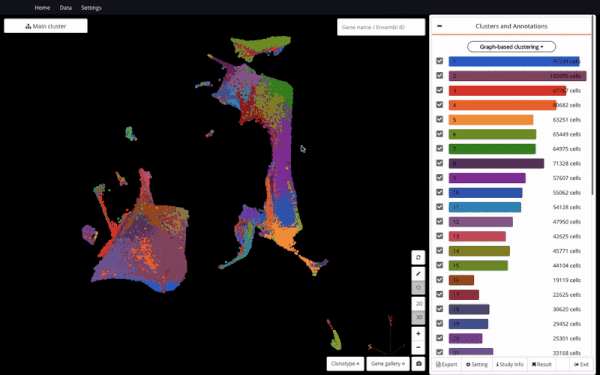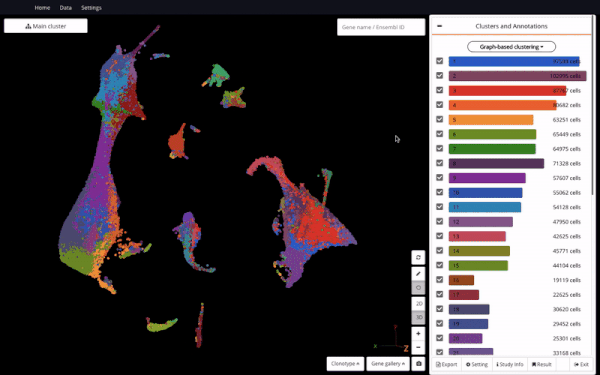
Mouse Organogenesis Cell Atlas (Cao et.al., 2019) in BioTuring Browser
BioTuring Browser is the first software to provide 3D rotation, differential expression analysis, and batch effect removal for many batches of scRNA-seq data. With the help of machine learning and AI, the software can run real-time cell type prediction, helping researchers and biomedical scientists quickly annotate cell populations. At the moment, our cell type prediction algorithm is optimized for neural cells and immune cells – the final aim is to cover all cell types in the body.
Here’s to name a few analyses integrated in BioTuring Browser:
- Transcript quantification with high accuracy and speed (using Hera-T)
- Integrate Seurat and Scanpy objects to the pipeline
- Batch effect removal with CCA, MNN, and Harmony correction
- Interactive 2- and 3-D visualization of up to 1.3 million cells
- Marker gene detection and real-time cell type prediction
- Sub-clustering
- Differential expression analysis
- Pseudotime lineage reconstruction
- Pairing clonotype data with single-cell expression data
- Analyzing cell surface data from CITE-Seq/ 10X Genomics Feature Barcoding technology
Together with a full range of analyses and visualizations, BioTuring Browser also provides scientists with easy access to latest scRNA-seq datasets from published studies, making it extremely easy to view and reanalyze valuable past works and compare such with their current studies. Above all, this library of published datasets promises to remove any barriers to access published data and make science completely transparent.
BioTuring Browser is now available for Windows, Mac, and Linux at www.bioturing.com/product/bbrowser. We also look forward to comments from users to improve the software, and if you need to adapt and customize BioTuring Browser for your company/organization, please don’t hesitate to contact us at: support@bioturing.com