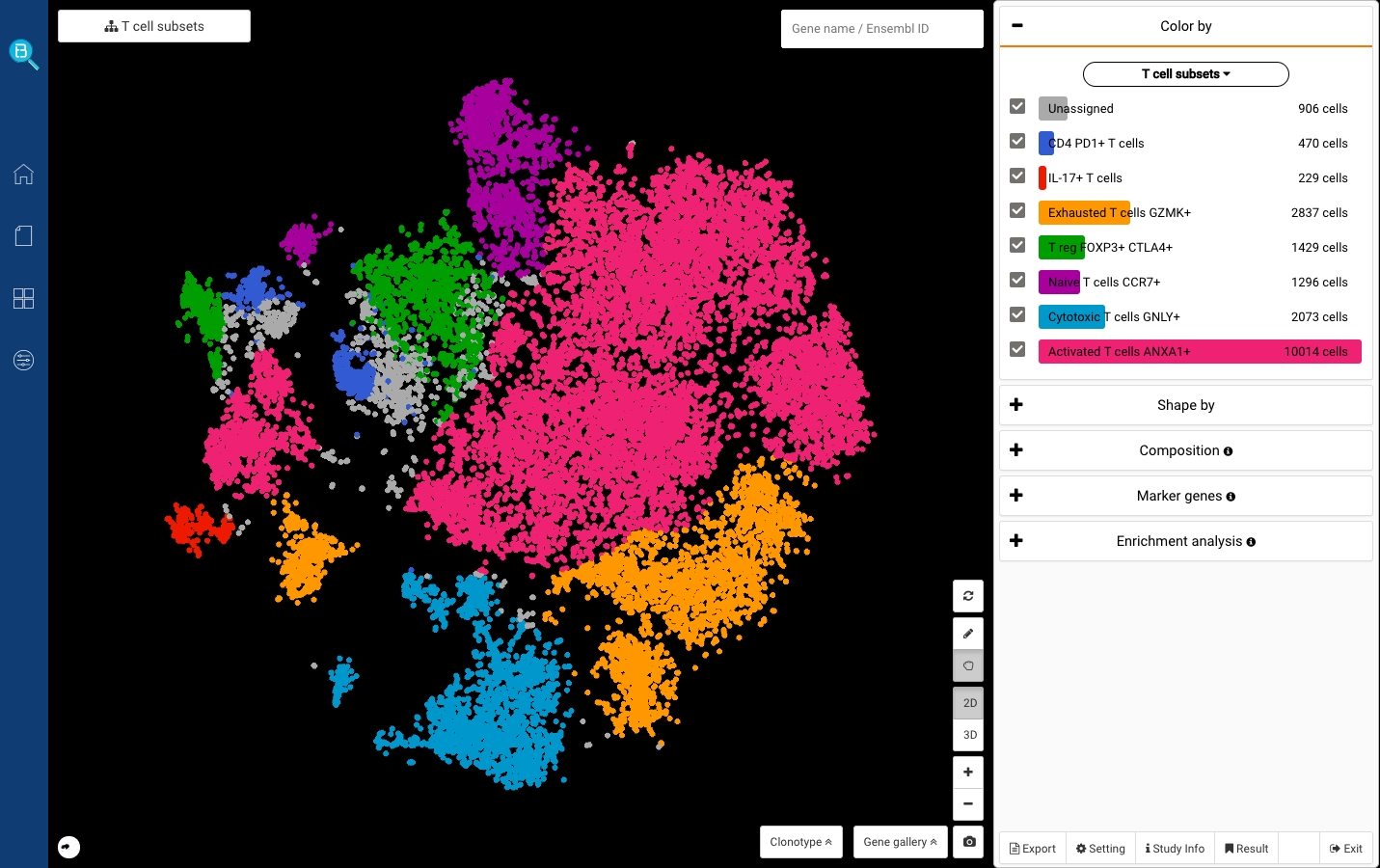
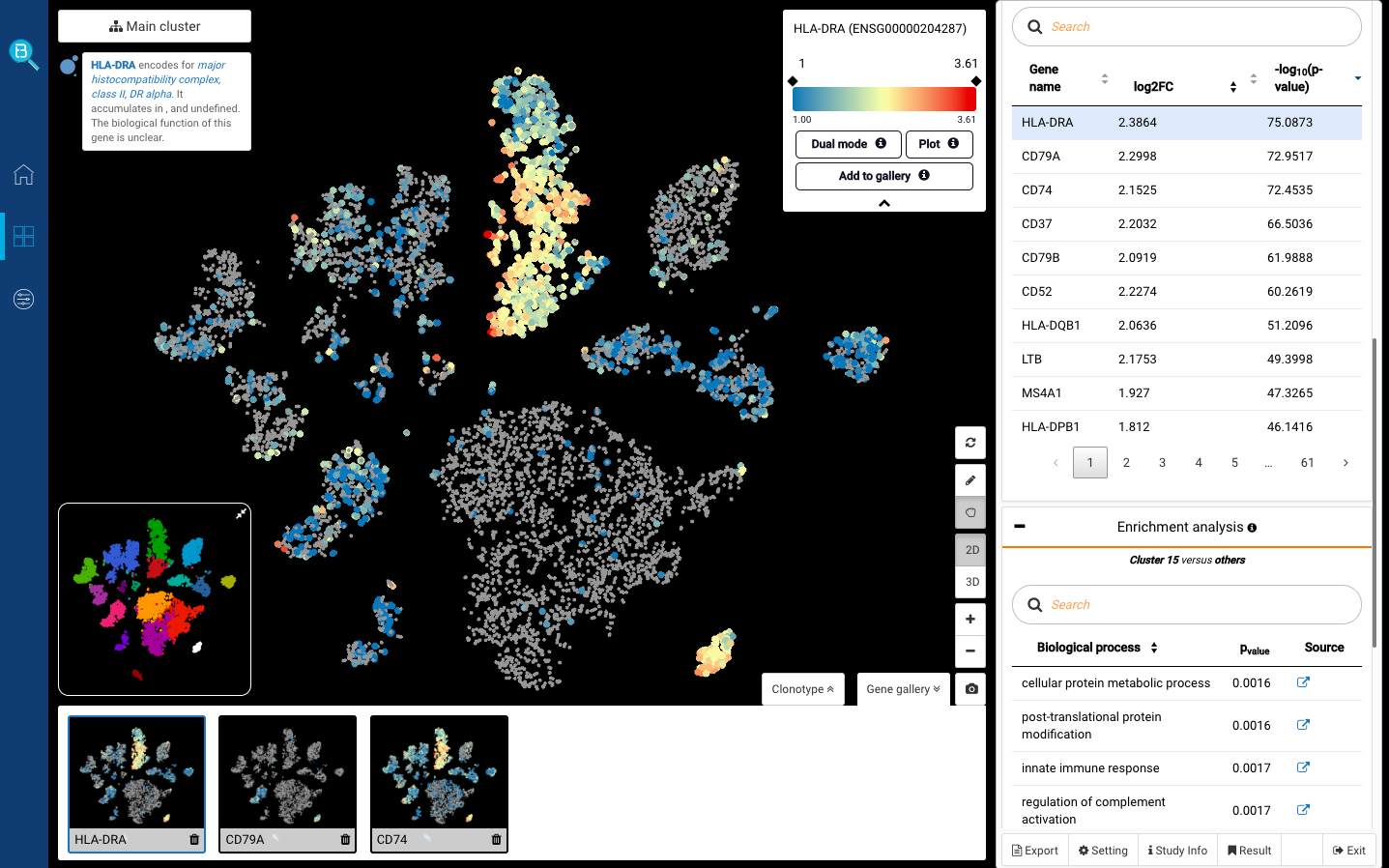
scRNA-Seq Cell Type Annotation: Common Approaches and Tools
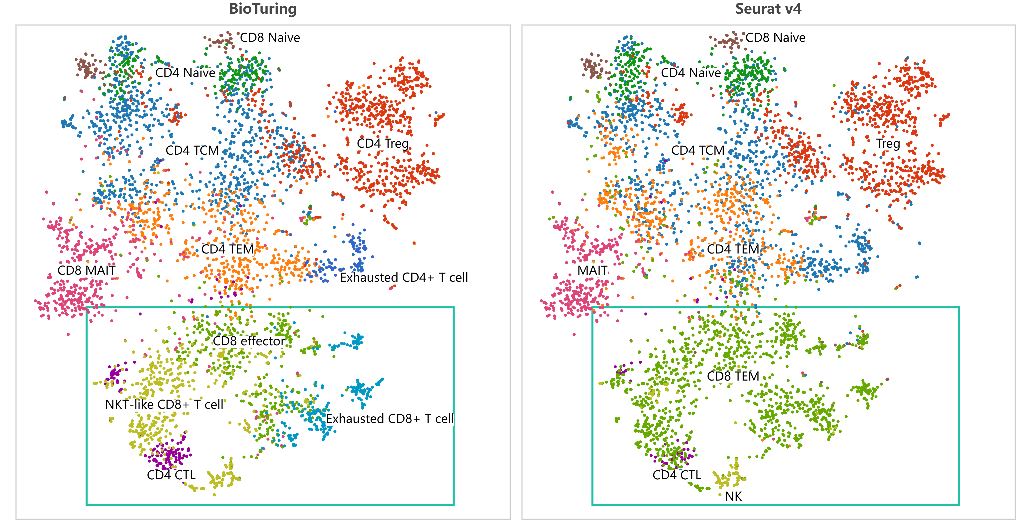
Assigning cell type identity to cells is a basic yet vital step required in single-cell RNA Sequencing data analysis (scRNA-Seq), often done after dimensionality reduction and scRNA-Seq clustering . If you have successfully captured informative clusters, it’s time to face an even harder challenge: identify what cell type or cell […]