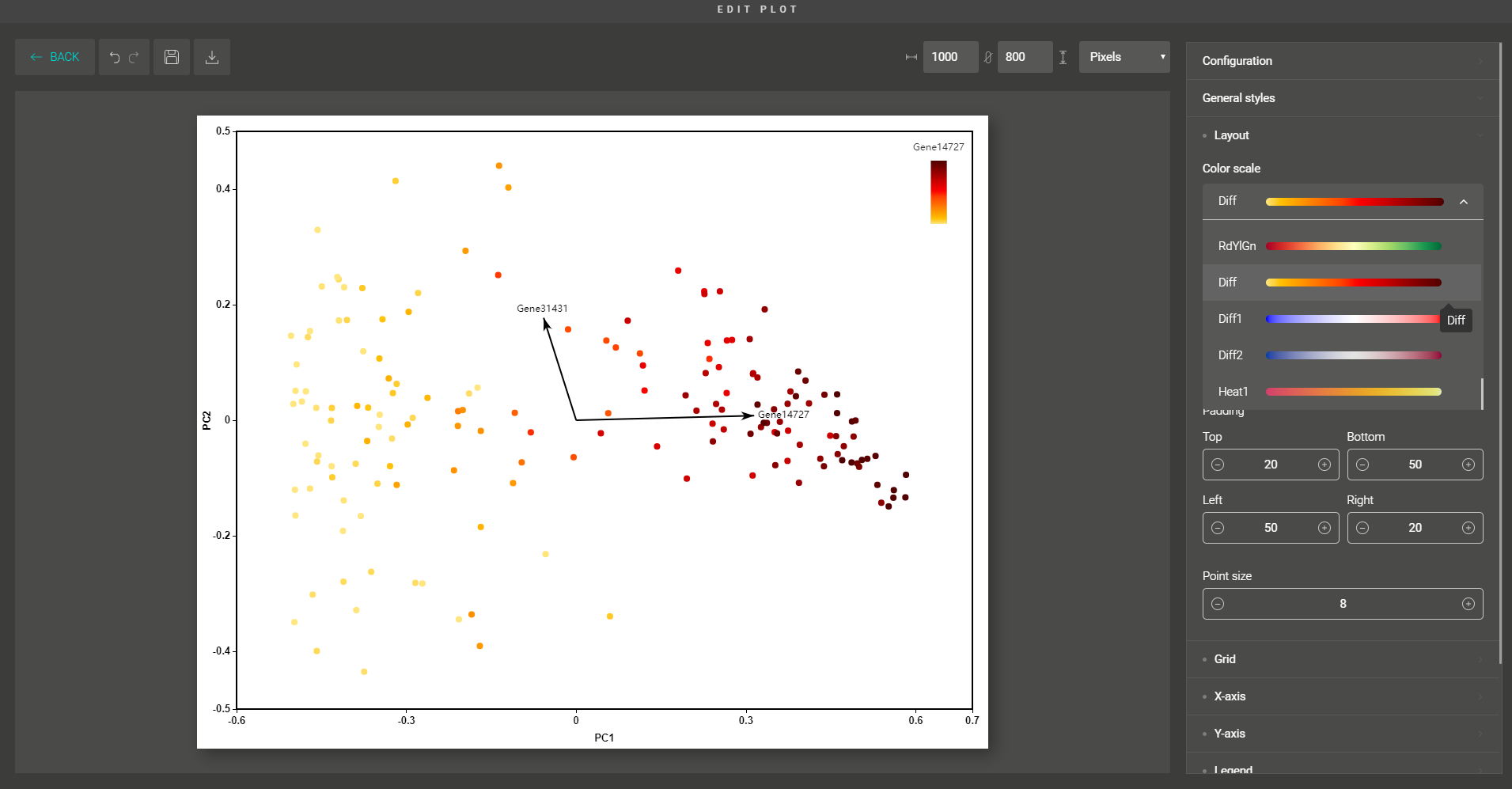
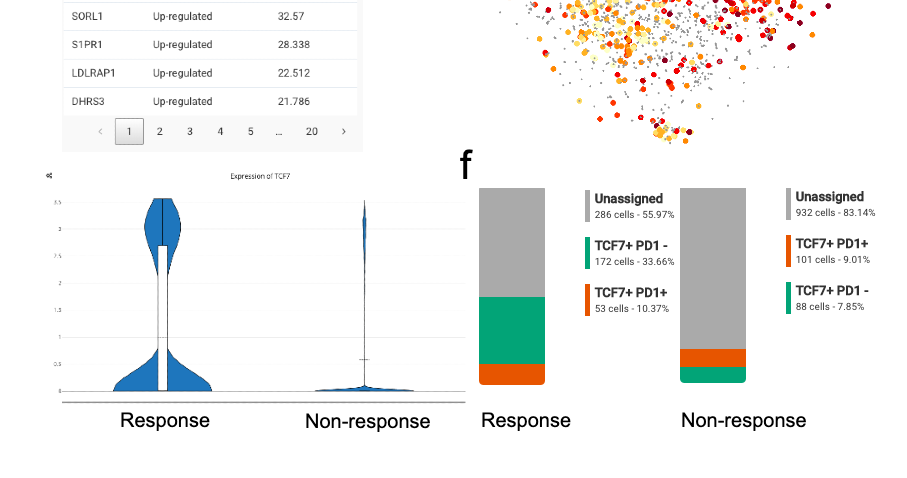
Zooming in the intratumoral heterogeneity of liver cancer with BBrowser single cell database
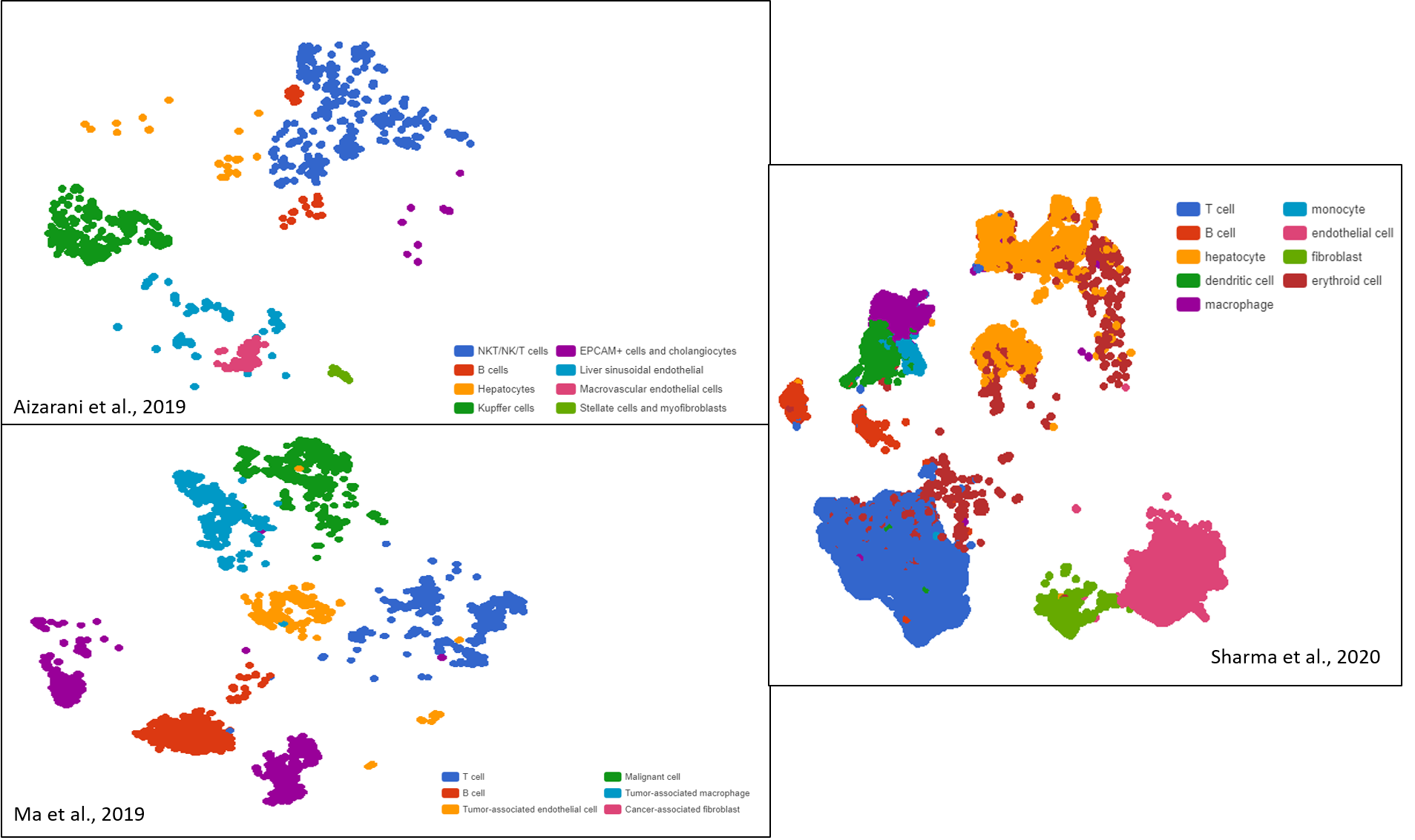
Introduction Why are tumors so resilient against available cancer therapies? One answer lies in intratumoral heterogeneity. Intratumoral heterogeneity describes the diversity in tumor cell populations. Cancer cells exhibit startling distinctions in their types, shape, metabolic activities, transcriptomic profiles, and more. The causes of intratumoral heterogeneity are both heritable (clonal expansion […]